You are here
Basic Science is the Best Protection Against Epidemics
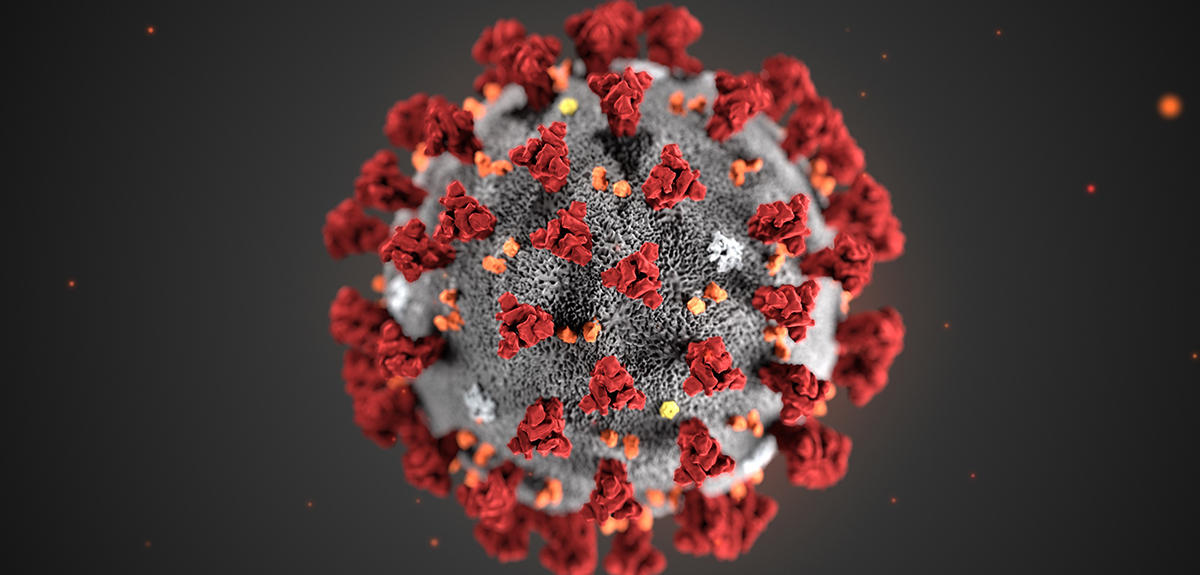
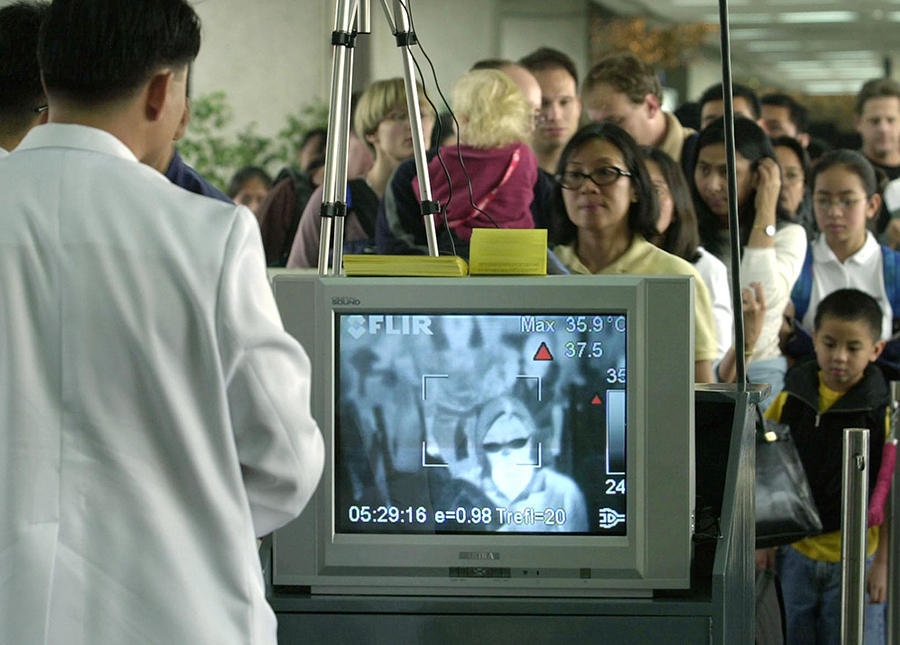
Since it first emerged in China at the end of 2019, the SARS-CoV-2 coronavirus has been spreading across the world, leading the WHO to designate the situation as a pandemic last week. First of all, what are your views on the emergence of this virus?
Bruno Canard:1 Our relationship with nature plays a key role in the development of this type of pathogen. Nature has been seen as a hive that one could pilfer without restraint…. until there are no more bees to make the honey. Global anthropic pressure has favoured the emergence of viruses that were hitherto hidden in animals and maintained in their natural habitats by high levels of biodiversity. Indeed, several studies have demonstrated that diversity is the best shield against the development of infections. The coronaviruses that caused Severe Acute Respiratory Syndrome (SARS), Middle East Respiratory Syndrome (MERS) and SARS-CoV-2 all came from the animal world and crossed the interspecies barrier.
How has knowledge improved since the start of the epidemic?
B. C.: We were not starting from scratch. Coronaviruses have been known since the 1950s but the scientific community only started to produce significant results following the advent of molecular biology in the 1990s. In my area, the first three-dimensional structure of a protein was obtained by Rolf Hilgenfeld in 2002. It went totally unnoticed as the virus (TGEV), which causes gastroenteritis in farmed pigs, was not popular with the media. On the other hand, his findings very rapidly enabled Rolf Hilgenfeld to establish the structure of the first protein, and principal protease, of the SARS virus, in 2004. As it happens, this type of enzyme (viral protease) is an interesting target for drug design. On our side, we generated the first original structure of a SARS protein a few months later, still in 2004.
One significant difference for the scientific community today is that it can now monitor an epidemic in real-time by high-throughput sequencing, in particular via the Nextstrain website, which is an open source project. It makes it possible to follow an individual who is infected in the field and within hours obtain the sequence of the virus in order to determine its phylogenesis. More generally, research teams are collaborating instantaneously to find a balance between disseminating knowledge – which is more essential than ever – and protecting their discoveries.
For example, following publication of the viral sequence by Chinese teams, scientists working with me – Étienne Decroly and Bruno Coutard (who is now at Aix-Marseille Université) – focused among other features on the specific dynamics of the virus. They detected a change in the Spike protein, which causes the contagiousness of the disease and its high capacity for transmission, compared with the 2003 infection. Our knowledge is improving, but there is much still to discover.
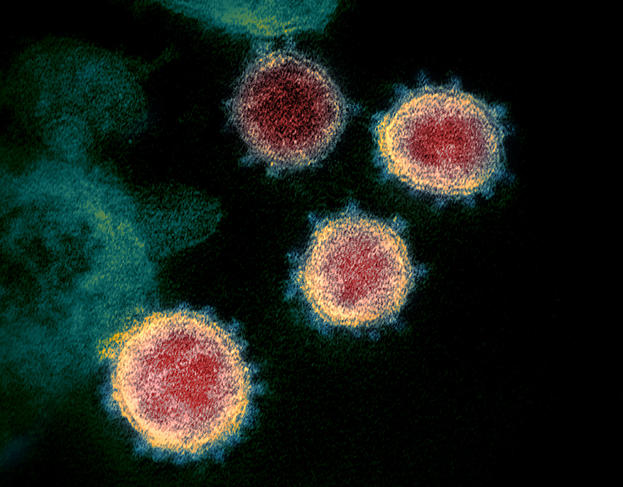
What other fundamental processes linked to coronaviruses are you studying as a priority at the moment?
B. C.: With Étienne Decroly, François Ferron and other members of the team, we are trying to understand how the RNA replication enzymes, namely methyltransferases and viral exonucleases, work together to evade the immune defences of the cell while “decorating” it with viral RNA. We shall be following the same path during the epidemic. My interest in replication enzymes originated from one of my teachers, Pierre Monsan, who said one day: “You see, polymerases are not like hydrolases: they choose a substrate out of thousands and manufacture highly complex substances at lightning speed. It’s more exciting than taking a water molecule and cutting a bond…”. I still subscribe to that view.
How useful can lessons from previous epidemics prove?
B. C.: There is a great deal to learn from the information and knowledge collected during previous epidemics involving other types of coronavirus. All this data is crucial because our most important battle at present is to determine the immunological behaviour of these viruses, understand what will make them shrink and find out whether some people are naturally immune, etc. How will the pathogen behave when it comes into contact with a new population that already has some immunity, or another population which has none at all? For example, from the data concerning patients affected by the SARS epidemic in 2003, one might wonder if the antibodies produced at that time might protect against the virus we are dealing with today.
What are the different therapeutic options that need to be explored?
B. C.: There are three main options: a vaccine, therapeutic treatments and the repositioning of existing compounds. At best, we need 18 months to create a vaccine, and years to produce a new drug. Repositioning, a solution that became popular in the early 2000s, has the advantage of being much more immediate.
The idea is to use compounds that have already been thoroughly screened and apply them to other pathologies. For coronaviruses, five drugs in particular are being used in clinical trials. Following the same principle, chloroquine, which has antiviral potential, has also been tested. However, it takes a large number of patients to validate the true effect of a molecule and obtain statistically conclusive results. Observing an improvement in the state of an affected individual is not sufficient to confirm that a treatment works: this result must be further confirmed via the usual scientific validation process of a publication by experts, so as to prevent potentially harmful consequences for sufferers. During the Ebola epidemic, drugs were repositioned without confirmed scientific foundations. Urgency is not conducive to sound decisions, which is the underlying problem of so-called compassionate treatments.
In the longer term, do you think it is wise to invest in a vaccine or new drugs?
B. C.: Vaccines are perfectly suited to dealing with known viruses. This has been proven by historical successes with yellow fever, measles or even influenza. We should nevertheless bear in mind that knowing a virus is not sufficient to find a vaccine against it, as has been the case for HIV or hepatitis C. Investing in a jab for coronavirus is a double challenge. That of knowing whether the infection will disappear or not (SARS-CoV lasted for six months in 2003) – and therefore taking the risk that the inoculation will become obsolete – and that of not being certain whether we will actually succeed in designing a vaccine. In this context, might it not be better to make more intelligent use of the sums allocated to developing a jab by investing in other therapeutic options?
Indeed, the two SARS viruses in 2003 and 2019 displayed almost perfect similarity in their replicative machinery. The replication enzymes of both (the targets for drugs) were the same because they were not seen to evolve, unlike the viral envelope (target for a vaccine) which is constantly attacked by the immunity of different hosts. Had drugs been developed as early as 2003 against these enzymes, they would be very effective in 2020 against the current virus, with no delay in their application.
The advantage of drugs over vaccines is that a single active substance is often sufficient to cover all members of a viral family. Such broad-spectrum antiviral agents would be more powerful because it would be enough to give the drug to one patient and to the cluster of individuals around them before the onset of symptoms. The virus would be killed immediately, thus eradicating the risk of an epidemic.

Why is this research path not generally preferred?
B. C.: The therapeutic solution has never been preferred since 2003 for several reasons. The first is cultural: there is a tradition of vaccines in France because of Louis Pasteur’s heritage, which encourages us to favour inoculation because it has proved its worth. The iconic image of Pasteur saving young Joseph Meister who had been attacked by a rabid dog still triggers a lot of emotion. Conversely, the fact that after an infection with smallpox an antiviral treatment works better than a vaccine does not have the same impact. And yet this finding was published in Nature in 2006. We must be careful, test other approaches and be aware that a jab is not always the best response to a virus.
Secondly, the search for new therapeutic treatments requires long-term investment. It uses high tech facilities and necessitates collaborations between disciplines that may range from structural biology to computing. The CNRS is in fact an organisation that is wholly adapted to conducting this type of work; it is its very purpose, its dedicated field, and its excellence, even though this expertise applies to the healthcare sector, which is also covered by the INSERM, Institut Pasteur and other highly competent actors. However, history has markedly reduced the opportunities for this type of research on coronaviruses. This domain suffered from the financial crisis in 2008 that led governments to redirect their financial support to other parts of society, and scientific policies (including the reform of tax credits the same year) which cut the budgets allocated to basic research.
How could such research help prepare to fight these viruses?
B. C.: Viruses occur suddenly but it is impossible to obtain immediate scientific results. The only solution is to anticipate what will happen. In 2015, with colleagues from Belgium and the Netherlands, we sent two letters of intent to the European Commission. We were targeting nine families of emerging viruses which required fundamental study. Since then, two have been the cause of epidemics: coronavirus and Zika. These are so many possibilities of an extremely contagious virus that is transmissible via the respiratory route (such as measles) occurring in the near future.
By anticipating and by studying the viral world in its broadest sense, it is possible to characterise a typical virus from each family, and in particular its mode of replication. In this way, a new pathogen will be similar to those we already know. In the event of an emergence, experiments already carried out could be used directly to obtain vaccines or antivirals, etc. The response would be immediate. Time is an investment for the future as regards this long-term research. Had we opted for this operational approach after the SARS epidemic in 2003, we would have saved considerable time in our search for appropriate drugs.
How is virological research organised?
B. C.: A lot of it is performed in networks and consortia that involve teams with complementary skills. Progress has been made concerning the sharing of reagents, which was hitherto not common practice when a virus was isolated. Now organisations such as EVA-GLOBAL (European Virus Archive – GLOBAL) distribute reagents and viruses to those approved laboratories that request them, and have greatly facilitated procedures. The nature of research has changed but in Europe it is constantly hampered by lack of funding. We have lost a great deal in terms of competitiveness compared with teams working in China.
During the 2000s, European research was based on large-scale projects and collaborative programmes. After the SARS outbreak in 2003, our team’s work enabled the description of several enzymes of interest for the design of new drugs, or simply helped elucidate how the virus was able to replicate so easily. In the late 2000s, virological research took a new turn; in both France and Europe, it shifted from anticipation to reaction.
Each epidemic pompts emergency funding that ultimately amounts to much less than that allocated in the 2000s to research aimed at anticipating such situations. Above all, epidemics are soon forgotten. A few years after the 2003 outbreak, political interest in SARS-CoV had already vanished.
What changes do you think are necessary for the future?
B. C.: At present, only a handful of laboratories are specialised in basic research on the replication enzymes and molecular drivers that will be the targets of future molecules against coronaviruses. Secondly, there has been some confusion between infective agents handled under high-security conditions and harmless recombinant proteins that fall under the same regulations, i.e. those concerning microorganisms and toxins (MOT). These lay down safety standards that are unsuited to basic science laboratories such as mine, and have somewhat sterilised all research. Things need to change in order to attract young talents, anticipate events intelligently and avoid the threat of epidemics. Sadly, I cannot see into the future. But what I can see is that serious, independent, rational and collegial research is our best defence against many outbreaks.
_________________________________________________________-
20 research projects focused on the epidemic
The Alliance nationale pour les sciences de la vie et de la santé (Aviesan)2 has mobilised its forces to accelerate research on the virus and on the CoVID-19 disease. Coordinated by the INSERM, the REACTing consortium has thus selected 20 scientific initiatives that target modelling of the epidemic, the search for treatment or preventive measures. As one of the project leaders with his colleague Étienne Decroly, Bruno Canard (see interview above) heads a programme on “Potentiating existing nucleoside therapies”.
More information on the projects selected is available on the website of the French Ministry of Higher Education, Research and Innovation.
- 1. CNRS research professor at the Laboratoire Architecture et Fonction des Macromolécules Biologiques (CNRS / Aix-Marseille Université).
- 2. Aviesan involves nine academic institutions that were founding members of the alliance: the CEA, CNRS, INRAE, INRIA, INSERM, Institut Pasteur, IRD, the Conference of University Presidents and the Conference of Directors General of Regional and University Hospital Centres.
Explore more
Author
After a degree in environmental studies at Paul Sabatier University in Toulouse, then in science journalism at Paris-Diderot University in Paris, Anaïs Culot worked in media relations at the CNRS and now collaborates with various magazines, including CNRS Le Journal, I'MTech and Science & Vie.